Who We Are
proteolutions is a Berlin-based biotech / bioinformatics service provider. We have developed a pioneering gene optimization algorithm that increases soluble protein yields in heterologous expression hosts far beyond the current state of the art. Unfavorable sequences are systematically avoided through our holistic approach that learns the optimal sequence neighborhoods from transcriptomic data and explores a large parameter space. Our optimized sequences will convey the highest reliability and best possible yields to your protein production endeavors in any host and at any scale.
One key parameter of all protein expression systems is the input DNA sequence, decisive for protein production, fold, solubility and lastly, high protein yields. Because of the combinatorial nature of codons, there is a plethora of possible sequences to choose from, even for relatively small proteins. For example, the redundancy of the genetic code allows for about 10200 different DNA sequences that can be synthesized to encode the human 387 amino acid protease pepsin. All of these sequences will yield different amounts of soluble target protein in given expression hosts. So it is a true challenge to find the optimal sequence for your target gene. See our Solutions overview for more information.

Our Method
To date, a multitude of strategies are available to locally adapt codon choices to the expression host's preferences, for example simple codon optimization, the codon adaptation index or the codon context. Analyzing these protocols, we find that despite an improvement compared to unoptimized sequences, they tend to be generally unreliable since they are restricted to local events in the gene sequence, disregarding connectivity – and this can have undesirable effects during gene expression.
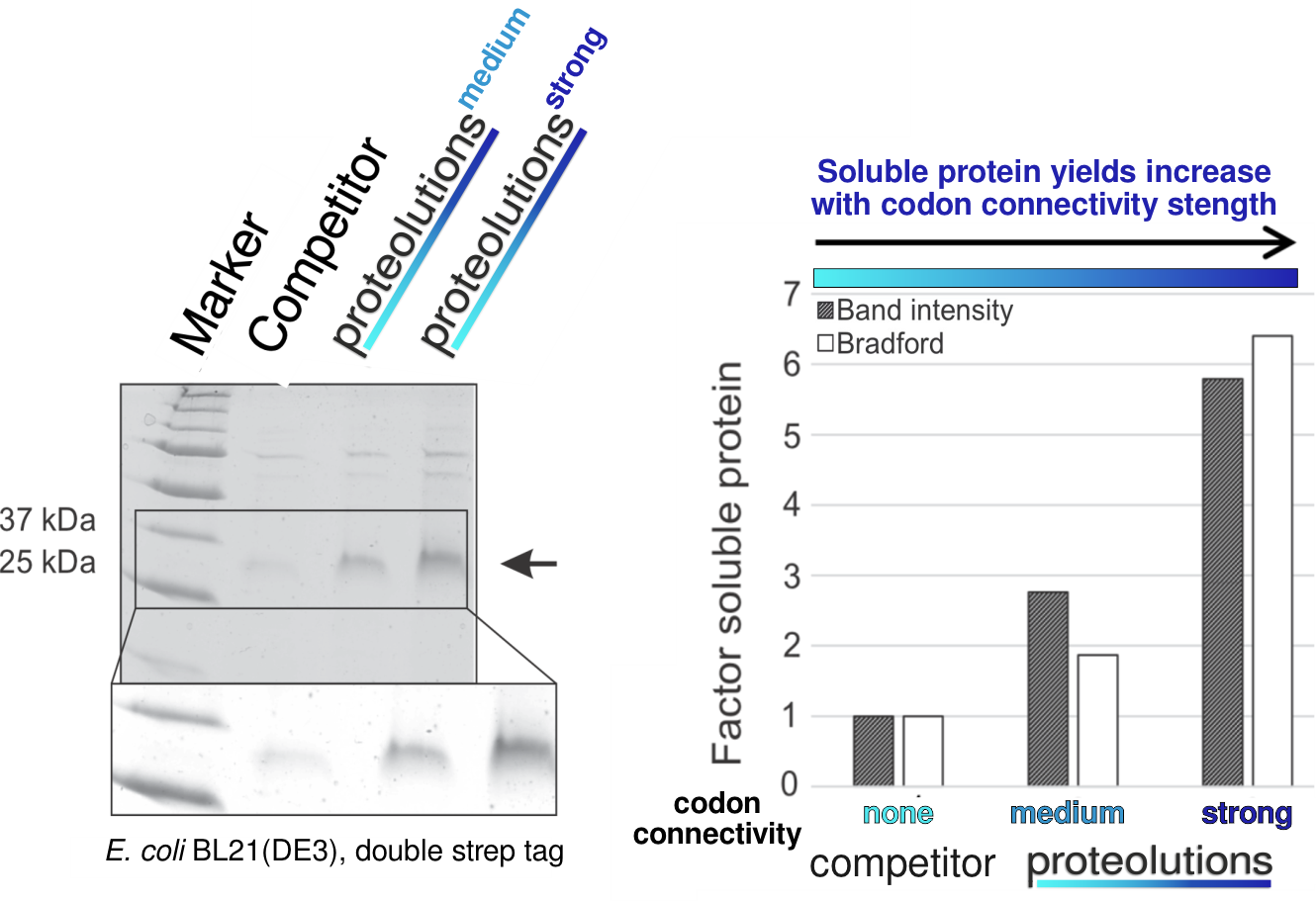
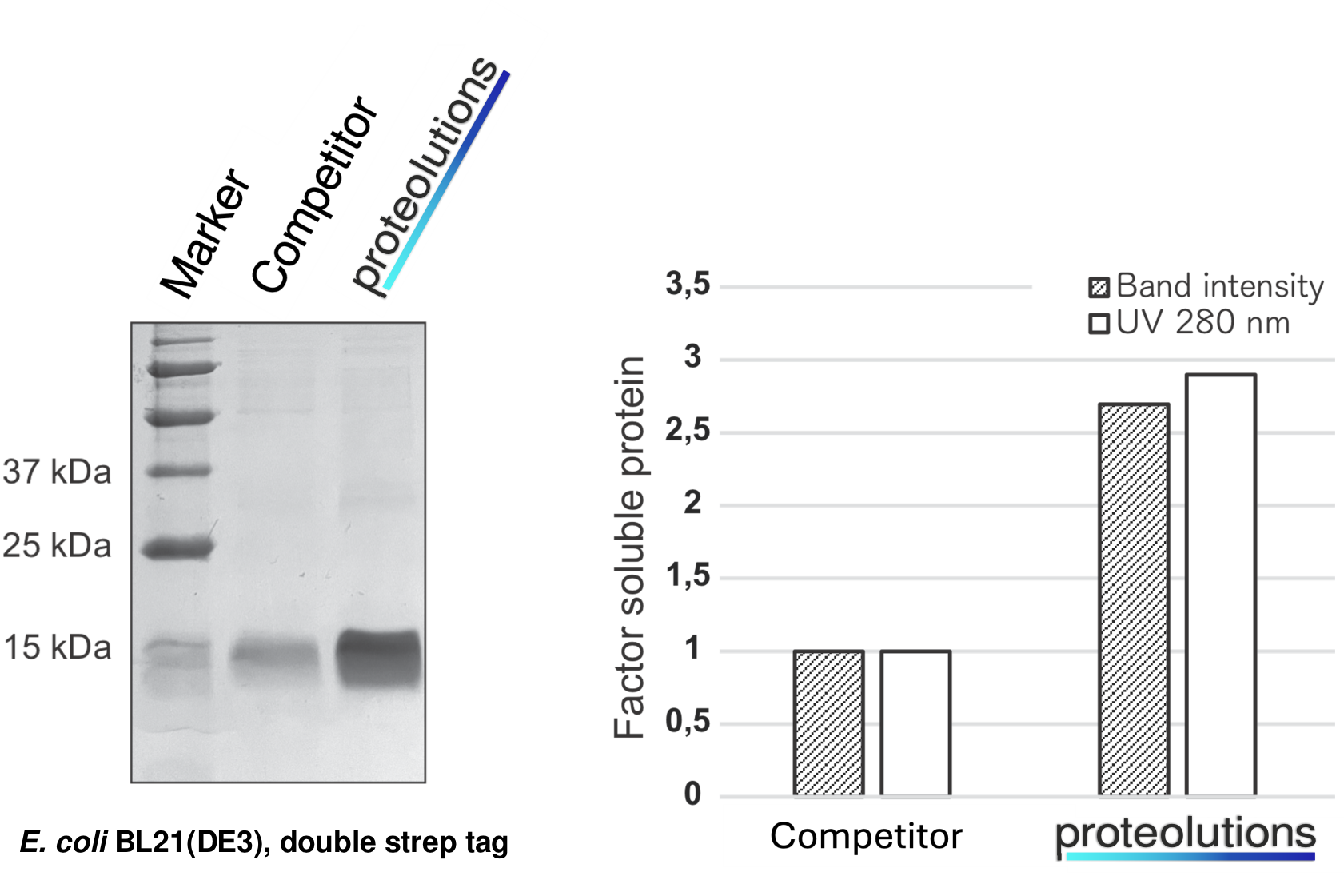
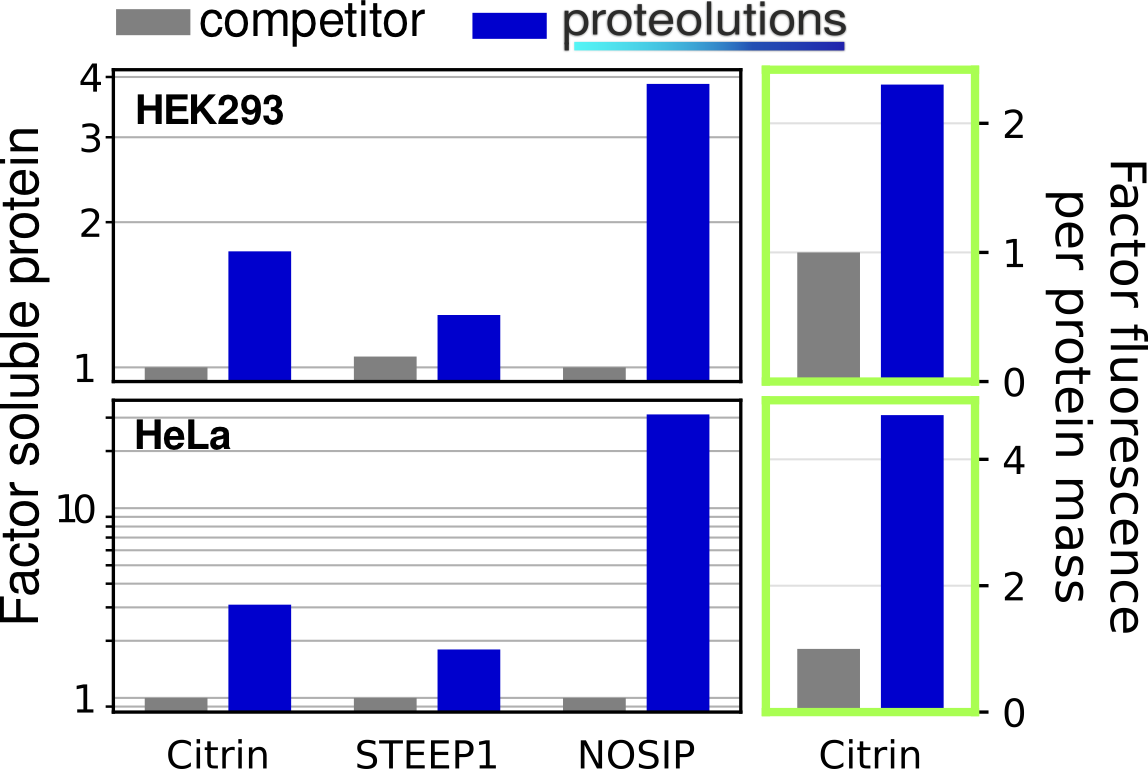
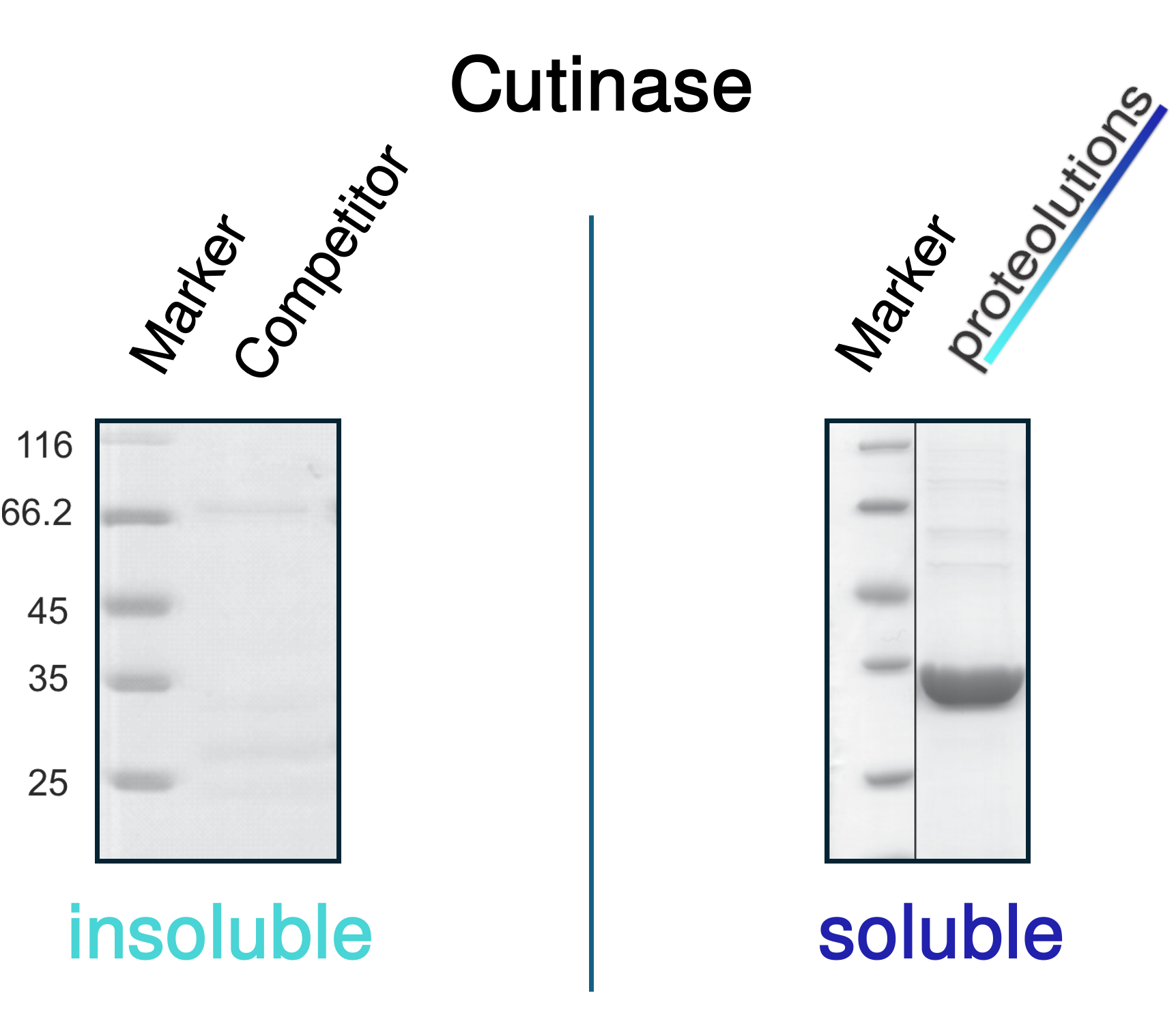
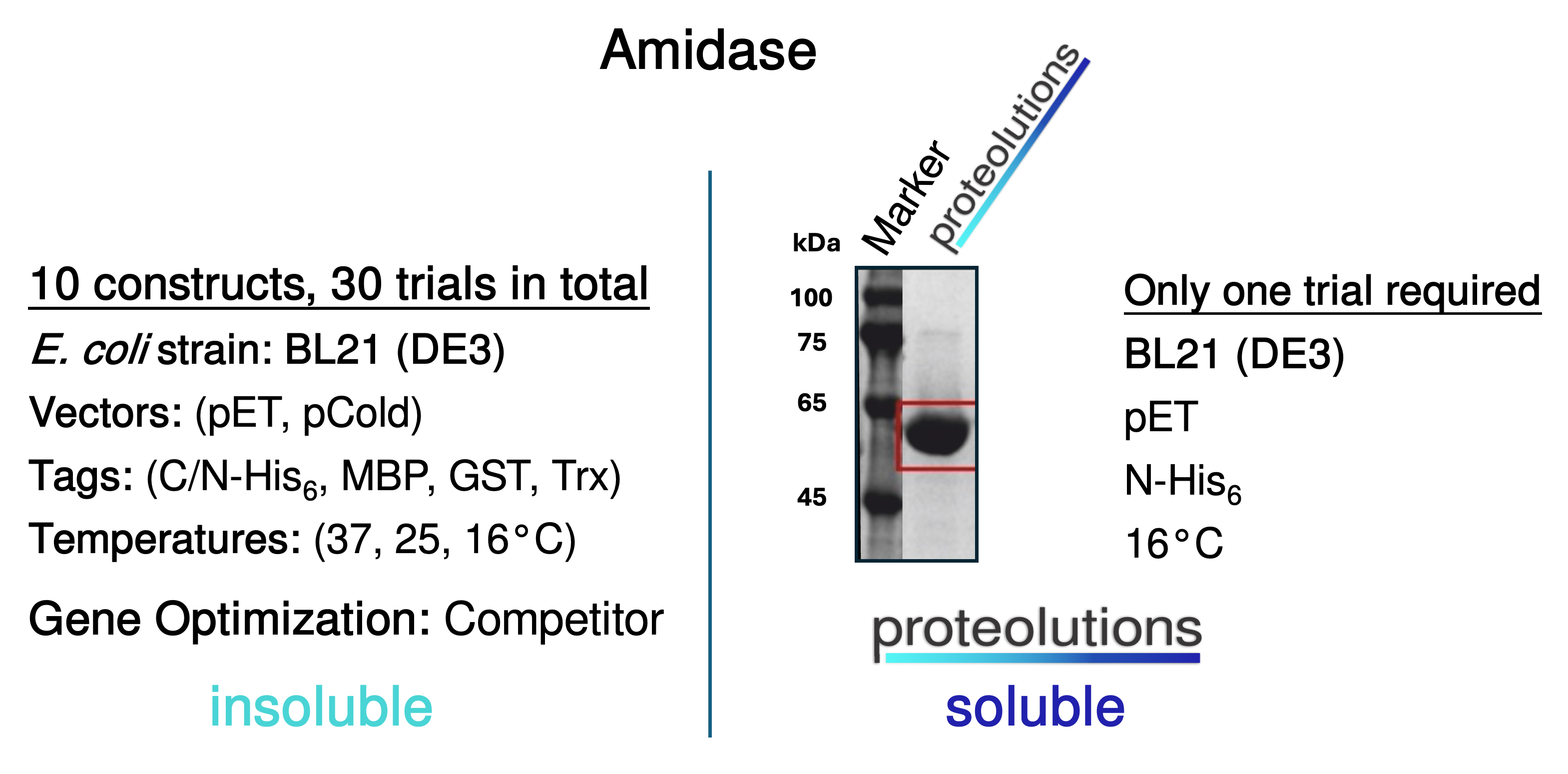
We at proteolutions have extensively tested our DNA optimization algorithm and the principle of codon connectivity. With the examples below we show that soluble gene expression directly increases with the strength of codon connectivity applied to the optimization and that our DNA optimization principle is universially applicable from bacterial up to mammalian hosts.